Overview and agenda¶
Overview¶
It may just be my imagination, but it seems like many practicing computational scientists have settled on the same basic workflow for highly repeatable computational analyses:
- a workflow engine, for long-running/compute-intensive analyses;
- some form of narrative documentation for exploring and tracking data analyses and graphing;
- a “cleanroom” execution environment for specifying and isolating dependencies;
- some form of version control for tracking digital artifacts and collaborating around them;
There are many different choices here, but some common ones are shown in the figure:
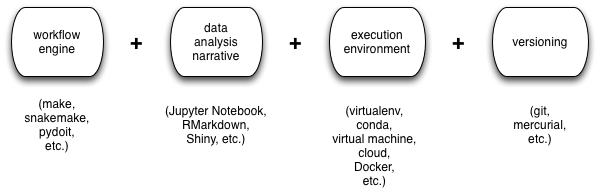
The way this all works in practice is shown in the next figure:
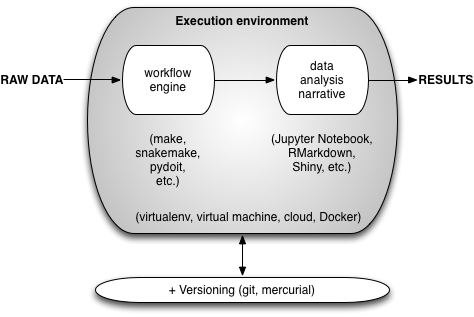
Basically, some form of virtual environment is used to execute the workflow and data analysis narrative, transforming raw data into results; all of the associated digital artifacts (except for the raw data and, generally, the results) are checked into some form of version control system.
What we will do today is build a small example that demonstrates all of these pieces working together.
Agenda¶
- Introducing Jupyter Notebook (CTB)
- (Re)visiting Make: a workflow engine (CTB)
- Lunch!
- Putting it all together: make, git, & jupyter notebook. (CTB)
- Docker for containerization. (CTB)
- Advanced topics (if time) (CTB)
- Travis: building things automagically (TKT)
LICENSE: This documentation and all textual/graphic site content is licensed under the Creative Commons - 0 License (CC0) -- fork @ github. Presentations (PPT/PDF) and PDFs are the property of their respective owners and are under the terms indicated within the presentation.